Congratulations to Michelle on her paper “Cdc42 GTPase regulates ESCRTs in nuclear envelope sealing and ER remodeling ” now published in JCB.
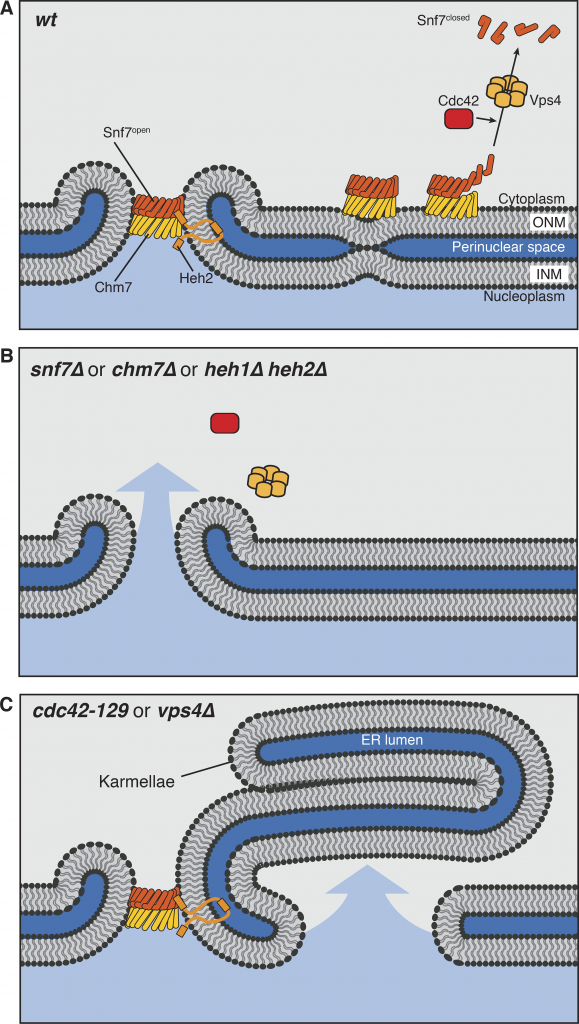
Cdc42, ESCRT-III, and Heh2 mutants share a leaky nucleus phenotype. (A) In normal cells, Heh2, Chm7, and Snf7 function at holes in the nuclear envelope to carry out annular fusion. ESCRT-III polymers are disassembled by Vps4, and we propose that Cdc42 is involved in the disassembly step by either directly contributing to Snf7 disassembly, activating Vps4 function, or cooperating with Vps4. (B) Mutants lacking components directly involved in annular fusion have holes in their nuclear envelopes left by nuclear fusion and ER fission events. (C) Cells lacking Vps4 and normal Cdc42 have unregulated ESCRT activity at the nuclear envelope. This causes the formation of nuclear karmallae and large holes in the nuclear envelope, leading to a defect in proper nucleo-cytoplasmic partitioning. ONM, outer nuclear membrane; INM, inner nuclear membrane.